TAU Study Finds Artificial Light at Night May Disrupt Biological Rhythms and Increase Mortality


Research
TAU Study Finds Artificial Light at Night May Disrupt Biological Rhythms and Increase Mortality

A new study from Tel Aviv University indicates for the first time that artificial lighting may disrupt natural rhythms of the immune system in wild rodents. According to the study, even exposure to minimal artificial light at night (ALAN), at intensities equivalent to standard street lighting, leads to a 2.35-fold increase in mortality.
The study was conducted at TAU's Zoological Garden, the I. Meier Segals Garden for Zoological Research on two local mammals, the golden spiny mouse and the common spiny mouse. It was carried out by doctoral student Hagar Vardi-Naim at the George S. Wise Faculty of Life Sciences. The study’s supervisors were Prof. Yariv Wine, head of the Applied Immunology Laboratory at the Shmunis School of Biomedicine and Cancer Research, and Prof. Noga Kronfeld-Schor, head of the Ecological and Evolutionary Physiology Laboratory at the School of Zoology, and Rector of TAU. Both Prof. Wine and Prof. Kronfeld-Schor are also affiliated with the new Environmental School at Tel Aviv University. The research was supported by the Israel Science Foundation. The disturbing findings were published in the journal Environmental Pollution.
Vardi-Naim explains: “Large parts of every mammal's body, including our own, are regulated by an internal biological clock. With a 24-hour rhythm based on the natural light-dark cycle, this biological clock signals to various organs and physiological systems, including the immune system, what they should do at different times of day. For example, the levels of certain white blood cells rise and fall in the blood, and the body produces more/less antibodies at specific times. Such oscillations can enhance the immune response to bacteria or viruses, but for this the body must know the time. Light pollution alters the natural light-dark regime, disrupts the central clock's synchronization with environmental time, and changes these patterns, rendering time almost meaningless.”
The researchers examined the effects of artificial lighting on the immune systems of two related species of small rodents: the golden spiny mouse and the common spiny mouse. Both live in the Israeli desert, sharing the same geographical habitat, but differing in their activity time: while the golden spiny mouse is active during the day, the common spiny mouse is active during night. The animals were taken from the Judean Desert to outdoor enclosures at TAU's Zoological Garden, where some of them were exposed to ALAN.
Vardi-Naim: “We kept the spiny mice in enclosures that simulated natural environmental conditions as much as possible. Half of the enclosures were illuminated at night with white LED, the most common type of lighting used today, at a relatively low intensity that simulates street lighting, while the control group was exposed only to natural light-dark conditions - the sun, moon, and stars.”
The researchers measured the percentage of white blood cells (i.e., lymphocytes) in the mice's blood at several points in the 24-hour cycle, and found a pattern similar to the human rhythm, with lymphocyte levels in the blood rising during rest hours, between two and four in the morning. In addition, they discovered a very clear 24-hour lymphocyte rhythm, and found that the amount of antibodies produced in response to an antigen (a substance that evokes the immune system's response, e.g. a virus or vaccine), is time-dependent.
“We saw that animals exposed to an antigen during their rest hours produced far more antibodies than those exposed during their active hours,” adds Vardi-Naim. “Exposure to light pollution, however, completely muddled these rhythms. Instead of a daily cycle of peaks and lows in the level of lymphocytes and immune response, we observed a complete flattening of the daily patterns. This means that the immune system loses its natural timing, and consequently, its response to infections, environmental stress, or vaccination might be less than optimal, possibly increasing the animals' vulnerability over time.”
In addition, extensive and rapid mortality was observed among the mice exposed to light pollution, with a 2.35 times higher risk of death compared to the control group. The researchers note that even though the exact cause of death could not be determined, the rise in mortality occurred alongside disruption of immune and endocrine (hormonal) rhythms, suggesting a likely connection between damage to biological timing and reduced survival.
Vardi-Naim emphasizes that the spiny mice in the study are only an example, and that the findings have implications for all living creatures, including humans. “Our results show that ALAN is not merely an aesthetic environmental change, but an active biological factor capable of disrupting critical physiological mechanisms. Chronic exposure to ALAN disrupted the timing of the mice's immune and endocrine systems and impaired their survival under conditions that otherwise simulated the natural environment. We believe that light pollution should be regarded as an environmental health risk with broad implications, not only for wildlife but also for human health and the ecosystem as a whole. Studies show that animals with weakened immune systems can transmit diseases to humans, and it is possible that the human immune system responds in a similar way. The study underlines the need to include biological considerations in lighting policies and to reexamine ALAN scope and intensity in both urban and open spaces.”
Overall, by studying animals that live in conditions close to their natural environment rather than in sterile laboratory settings, this research highlights the value of using wild models to understand how the immune system functions in the real world. Such approaches reveal how environmental changes, including growing light pollution, can affect complex biological systems in ways that are often missed in traditional lab studies. As human activity continues to reshape natural environments, studying immune responses under realistic ecological conditions is essential for understanding how global environmental change may influence the health of wildlife, ecosystems, and potentially humans.

Research
A TAU-led team are reviving lost popular Yiddish plays, starting with the TAU premiere of an original Hebrew adaptation of a buried classic

At Tel Aviv University’s theatre department, imposing ghostly white figures sing to an old woman on a stage designed like a surreal graveyard. This is the recent production of The Dybbuk, a comedic Yiddish play not performed since the turn of the century. The play is part of a sweeping inter-institutional project to restore popular Yiddish performance pieces and return them to the stage, led by TAU’s Dr. Ruthie Abeliovich. By bringing these plays into the 21st century, the researchers reveal truths about Jewish history and shed light on the critical cultural discussions that our modern Jewish community is still having today.
Laughter Before the Tragedy
Two lovers, an arranged marriage, and a demonic possession: many in the global Jewish community have heard of The Dybbuk, a Yiddish ghost story adapted to a famous play and even to a well-known kids’ movie (The Corpse Bride, Tim Burton). The play, by S. An-sky, is a tragedy about a doomed marriage and is considered an important piece of classical theatre. But not many know that decades before its creation, a comedy of the same title and themes, most notably made famous by playwright Joseph Latainer, swept through the Yiddish-speaking theatre world reaching millions of Jews.
Likely because of their mass appeal to common audiences, it and many other popular Yiddish shows were delegitimized by intellectuals as “shund” —trash—and effectively erased from the cannon. The devaluation of popular cultural art is a phenomenon we still see today, despite its broad influence.
“Studying these forgotten plays from the flourishing turn-of-the-century Yiddish theatre scene offers insight into the values, challenges and attitudes of the Jewish community at the time--and today."
Now, in a years-long project supported by a prestigious grant from the European Research Council, an international team of Yiddishists led by TAU Dr. Ruthie Abeliovitch of the Katz Faculty of Arts is resurrecting these forgotten dramas. “Studying these forgotten plays from the flourishing turn-of-the-century Yiddish theatre scene offers insight into the values, challenges and attitudes of the Jewish community at the time,” she says. “These shows were often developed in the New York Yiddish community and made their way to Europe. Because they were so popular, they give a window into mainstream mentalities all over the pre-Holocaust Jewish world—as well as revealing how we Jews today are still dealing with many of the same communal issues.”
Instead of tragedy, Latainer's show ends in a wedding. (Photo: Tami Shaham)
The research, fittingly dubbed “DYBBUK”, encompasses many different facets. The researchers have digitized hundreds of pages of handwritten performance notes through a newly-developed AI interface; begun creating a database of scripts for public use; and mounted an innovative Hebrew-language performance of The Dybbuk through TAU’s Department of Theatre Arts and the Buchmann-Mehta School of Music.
100 Years Later, The Dybbuk Hits the Stage
Latainer’s The Dybbuk: in the Clutches of Fanaticism follows Amelia, a young Jewish woman in love with her French teacher, a secular Jew named Leon. Her religious grandmother, however, has promised her to the local rabbi in exchange for his blessings. With assistance from Leon’s servant, Falik, the couple makes a desperate attempt to elope but is soon apprehended by the rabbi and his followers. In the end, the rabbi is exposed as a fraud, and Amelia and Leon get married.
The show deals with themes that follow us to this day: tradition vs. modernization; duty to community vs. individual fulfillment; the gendered politics of marriage, and more.
Drawing from a number of different accounts of productions, TAU staged a truly one-of-a-kind musical theatre performance. In translating and adapting to modern Hebrew, the team aimed to incorporate Yiddish as a living language and culture. This meant designing sets and costumes, composing music, adapting scripts and directing actors all in a way that would be recognizable and entertaining to audiences both 100 years ago and today. Many of the artists and all of the performers were TAU theatre and music students; the department also worked with Yonatan Levy, a professional director, Noam Enbar, a musical composer, and Shirly Marom, the project producer.
In a dream, the ghost of Amelia's mother tells her grandmother that she must let Amelia marry as she wishes. (Photo: Tami Shaham)
Going Digital
To recreate the show for adaptation, the researchers cross-referenced pages upon pages of notebooks recounting multiple different productions and versions of The Dybbuk. Since many were from over 100 years ago and not always well-preserved, these prompting notebooks—containing scripts, stage directions, musical cues, and annotations—presented a challenge. The team met that challenge with a modern solution: they created a new AI tool. Though handwriting recognition software is well established, no such deciphering model existed for Yiddish handwriting, especially on weathered pages.
“The plan is to make Yiddish plays, music, and art publicly available, so that these once-beloved works can live again."
Through detailed, painstaking work, the team trained a new machine-learning model that could digitize the prompting notebooks. To date, they have digitized over 700 pages of notes and made their software available to other Yiddishists for other restoration projects. Their data has also become the basis for the development of other Yiddish handwriting recognition programs. The digital project was led by Dr. Sinai Rusinek.
In addition to The Dybbuk, nearly 15 other plays have been transcribed and uploaded to an open database so that anyone can access and perform them. “The plan is to continue growing the database and making plays, music, historical insights, recordings and other aspects of Yiddish art publicly available,” says Dr. Abeliovitch, “so that these once-beloved works can live again.”

Research
TAU researchers develop ultra-efficient graphene switch at the nanometer scale

A team of researchers from Tel Aviv University, in collaboration with colleagues from Japan, has taken an important step toward the next generation of electronics. The scientists achieved highly precise control of the internal structure of graphene — an exceptionally thin and strong material — using a minute, nearly negligible amount of energy.
The study was conducted under the supervision of Prof. Moshe Ben-Shalom of the School of Physics and Astronomy, together with Prof. Michael Urbakh and Prof. Oded Hod of the School of Chemistry. The experiments and calculations were led by Dr. Nirmal Roy and Dr. Pengua Ying, supported by Simon Salleh Atri, Yoav Sharaby, Noam Raab, and Dr. Youngki Yao. The findings were published in the journal Nature Nanotechnology.
Graphene, which consists of a thin layer of carbon atoms, has long been regarded as a “star” in the world of materials. Yet it is not only the material itself that matters, but also how the graphene layers are stacked on top of one another. Different stacking arrangements create entirely different properties: different electrical conductivity, different responses to magnetic fields, and even conditions that enable the emergence of superconductivity.
Until now, controlled switching between these stacking arrangements has been a complex process that required a great deal of energy and was unsuitable for practical applications. In the new study, the researchers succeeded in overcoming this obstacle.
The solution they developed is based on an elegant concept: creating tiny “islands” of graphene — only tens of nanometers in diameter — where the layers remain in direct contact with one another, while the surrounding areas are separated by a layer that allows nearly frictionless sliding. Within these islands, one graphene layer can be shifted relative to another, thereby changing the stacking arrangement.
The result is striking: the material’s state can be changed using an extremely small force, with an energy input orders of magnitude lower than that required by existing memory technologies. In many cases, once the change is initiated, it continues on its own, without the need to apply additional force.
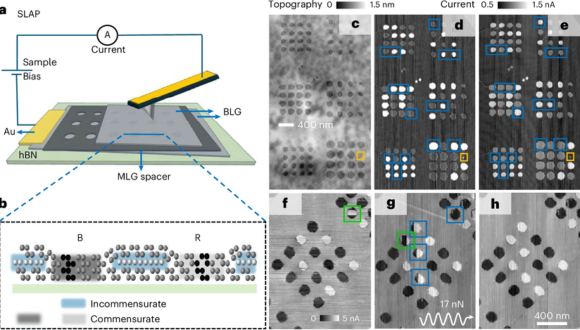
An illustration of the research: The Super-Lubric Array of Polytypes (SLAP) device in action. The bright and dark circles represent high and low electrical currents.
Beyond this, the researchers showed that neighboring islands can be connected so that a structural change in one island also affects its neighbors. This opens the door to creating systems in which different regions “communicate” with one another in a mechanical-elastic manner, similar to a neural network. Such a property may be particularly relevant to the development of neuromorphic computing — computers that mimic the way the brain operates.
According to the researchers, the new method opens promising avenues for the development of memory components, sensors, and tiny electronic devices that are both fast and exceptionally energy-efficient. In the future, it may enable the creation of smart electronic systems on the nanometer scale — systems that consume less energy, generate less heat, and can perform complex operations in ways that until now seemed purely theoretical.
Prof. Moshe Ben-Shalom concludes: “This is a breakthrough that has the potential to transform the way electronic components are designed at the nanometer scale. We show that it is possible to control the structure of graphene and other layered crystals in a precise, reversible, and extremely energy-efficient manner. Instead of breaking and rebuilding chemical bonds, we simply slide atomic layers over one another — a natural process that is much faster and more efficient. The ability to design interactions between different regions within a material opens up new possibilities, not only for advanced electronics but also for brain-inspired computing systems. This is another step toward turning physical phenomena that until now were considered purely academic into practical, working technology.”

Research
Rare archaeological findings in the Sakhnin Valley suggest that Homo erectus attributed meaning and visual significance to natural features hundreds of thousands of years ago

A Tel Aviv University Archaeologist and a resident of the Arab city of Sakhnin recently led an exceptional archaeological discovery in Lower Galilee Sakhnin Valley, shedding new light on the cultural and cognitive world of our early ancestors. A surface survey revealed a series of Paleolithic sites containing hundreds of handaxes — large, carefully crafted stone tools — identified with Homo erectus, the early human species that lived in the region hundreds of thousands of years ago.
Beyond the impressive quantity, however, the most noteworthy and unique find is an unprecedented concentration of handaxes shaped deliberately around fossils and distinctive geological features — a phenomenon almost unknown from other sites around the world. The study appeared in the prestigious journal published by the Sonia & Marco Nadler Institute of Archaeology, Entin Faculty of Humanities: Tel Aviv: Journal of the Institute of Archaeology of Tel Aviv University.
The sites in the Sakhnin Valley were identified by Muataz Shalata, a self-taught nature enthusiast from the city of Sakhnin, who noticed unusual knapped stones scattered across the terrain. He contacted Prof. Ran Barkai of the Elkov Department of Archaeology at Tel Aviv University, an expert in the study of early Paleolithic cultures. Together, they are leading an innovative study focusing on human behaviors that evolved in the Sakhnin Valley hundreds of thousands of years ago.
Prof. Barkai: “Handaxes served as the main tool of early humans for more than a million years, and are known from Africa, Asia, and Europe. In the Sakhnin Valley, many hundreds of handaxes were found, indicating that the area served as an important hub of human activity over long periods of time. Providing early humans with all their needs - water sources, game, and an exceptional abundance of high-quality flint nodules — the area probably attracted human groups repeatedly over hundreds of thousands of years”.
"The Valley is also very rich in geodes - rounded, brain-like geological concretions containing sparkling crystals, as well as flint nodules embedded with fossil remains. Early humans who came here hundreds of thousands of years ago must have been astonished by this exceptional richness of stones, leaving behind them an extraordinary phenomenon: we have discovered more than ten handaxes fashioned from flint nodules containing fossils or special geological formations, with these natural features deliberately preserved in a prominent position at the center of each handaxe. Since such features make precise and symmetrical knapping difficult, we can conclude that the selection of these specific stones was not accidental. On the contrary — the knapping process highlighted the natural feature and kept it at the center of the tool."

A handaxe shaped around the imprint of a fossil from the Sakhnin Valley
According to the researchers, this unique phenomenon clearly demonstrates aesthetic and conceptual intention among early humans, beyond functional considerations of tool production. Embedded fossils and geological formations do not improve the tool’s performance and may even impair it, yet such stones were repeatedly preferred as raw material. The conscious choice to invest effort in shaping a tool around an exceptional natural feature indicates that beyond survival needs, humans attributed special value to the stones' appearance and meaning. Knapping served as a means for framing, highlighting, and enhancing intriguing natural phenomena, reflecting advanced perceptual and cognitive abilities.
The researchers also note that the Sakhnin Valley is located near presumed routes of early elephants, which were a primary food source for humans during this period. Thus, as at other prehistoric sites such as Gesher Bnot Ya‘akov, the handaxes were probably used to cut up elephants and extract calories from their fat and meat. However, such a high concentration of special handaxes is unknown in any other site worldwide, exceeding all comparable finds documented to date.
Prof. Barkai concludes: “The unique landscape of the Sakhnin Valley led early humans to behave in a distinctive manner. Apparently, they attributed great significance to the fossils and special geological features they found in the Valley, regarding them as manifestations of the potency, primordiality, and wonder of the cosmos. The integration of fossils and geological features endowed the handaxes with added potency and meaning, connecting them primeval elements. The findings from the Sakhnin Valley open a rare window into the inner world of early humans, indicating that already at the dawn of human history they were sensitive to aesthetics, attributed meaning to nature, and had complex relationships with their world. The discovery places the Sakhnin Valley and the Lower Galilee at the heart of the international scholarly discussion on the origins of cognition, aesthetics and meaning in human life.”
* Prof. Ran Barkai is a prehistoric archaeologist at Tel Aviv University, specializing in the study of early human evolution and cultures. He is known, among other things, for his excavations at Qesem Cave and his research on the links between diet, animals, and the development of human consciousness.